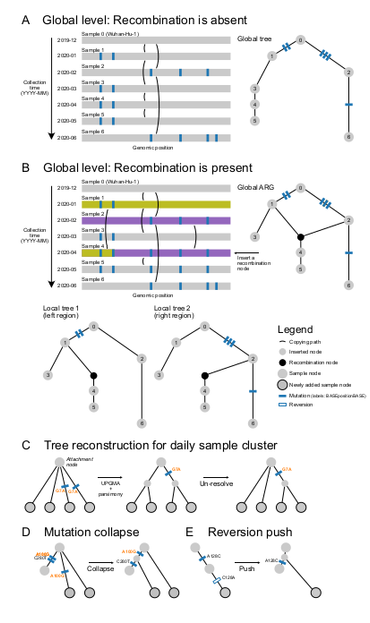
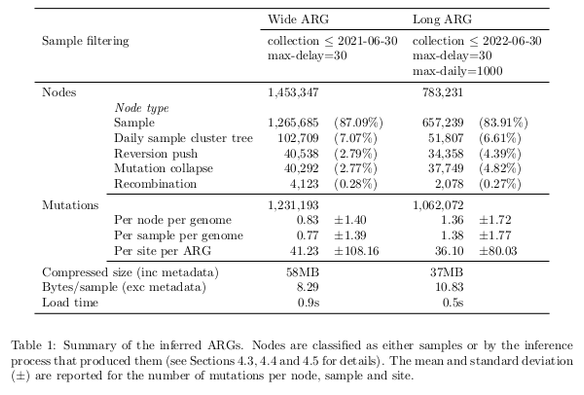
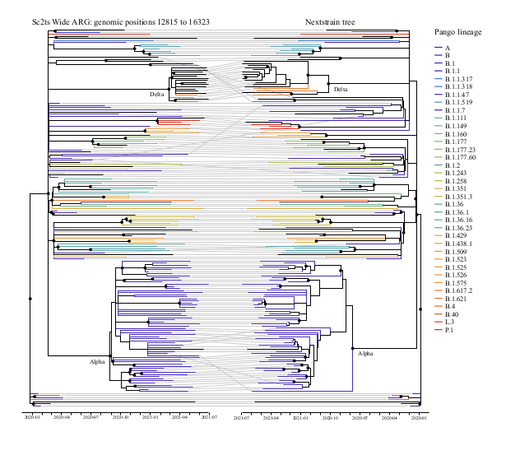
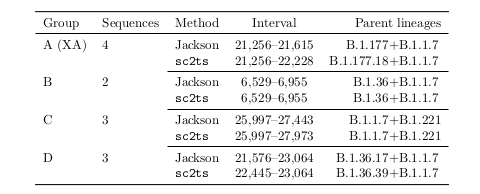
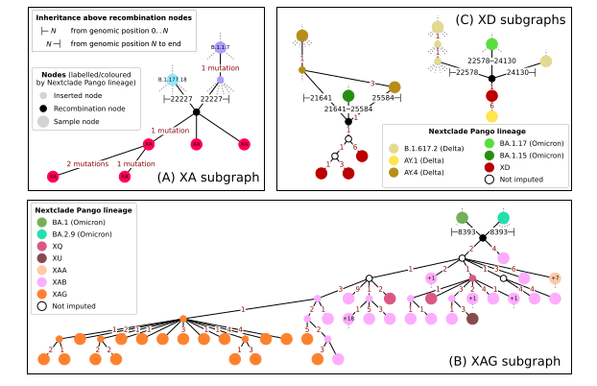
Tootorial on the recent preprint on inferring ARGs for SARS-Cov-2:
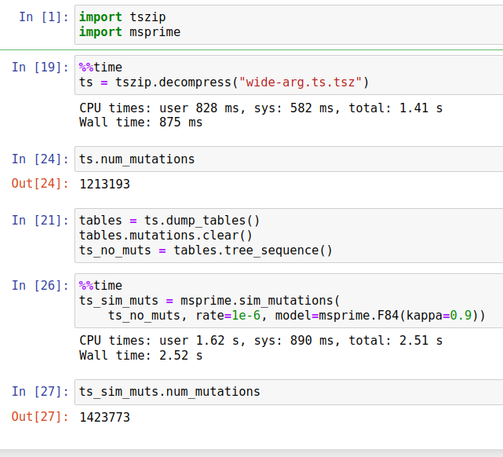
We can analyse these ARGs using the well-established and feature-rich #tskit library (and surrounding ecosystem). All of the analyses for the preprint were done using Jupyter notebooks, and most run in seconds on a standard laptop.
The sc2ts method itself is a thin wrapper around existing high performance components. The actual inference code is only about 1300 lines of Python. We can infer viral ARGs at this unprecedented scale with no compiled code **at all**.
https://github.com/jeromekelleher/sc2ts/blob/main/sc2ts/inference.py
ps. See also the summary post on virological:
https://virological.org/t/towards-pandemic-scale-ancestral-recombination-graphs-of-sars-cov-2/936
Towards Pandemic-Scale Ancestral Recombination Graphs of SARS-CoV-2
Towards Pandemic-Scale Ancestral Recombination Graphs of SARS-CoV-2 Shing H. Zhan1, Anastasia Ignatieva2,3,*, Yan Wong1,*, Katherine Eaton4, Benjamin Jeffery1, Duncan S. Palmer1, Carmen L. Murall4, Sarah P. Otto5, Jerome Kelleher1,† 1Big Data Institute , Li Ka Shing Centre for Health Information and Discovery, University of Oxford, United Kingdom 2Department of Statistics, University of Oxford, United Kingdom 3School of Mathematics and Statistics, University of Glasgow, United Kingdom 4Nati...