Proud and sad in equal measures to see I didn't contribute any code to the latest #msprime release. It's all grown up now 😢
🌲 New #msprime release: 1.3.0 🌲
This release introduces:
1/ additional nodes API: a new flexible way to store information on the ancestors associated with a custom subset of all possible events that might happen in the history of a sample.
2/ the simulation of microsatellite repeats.
Check out the details here: https://tskit.dev/news/20231225-msprime-1.3.0.html
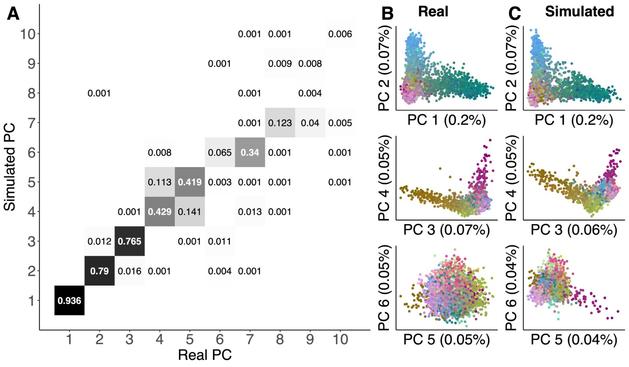
Ultra-realistic simulations of 1.4 million human genomes generated by using a detailed pedigree of French Canadians as input to #msprime! These simulations are hugely useful for large-scale genomics methods development, because they are freely available, easy to download (2.8G for chr1), and efficient to process using #tskit. I hope they will become a standard benchmark across all sorts of methods.
Here's the repo on GitHub for the ARG visualizer which plots a #tskit tree sequence using D3.js:
https://github.com/kitchensjn/tskit_arg_visualizer
You will need #jupyter, #msprime, and #numpy installed in Python to run the notebook. Let me know if you have any issues getting it running.
There are so many rules around node positions and paths between nodes. I'm slowly picking through a document of scenarios that break the current implementation. More updates to come!
GitHub - kitchensjn/tskit_arg_visualizer: Interactive visualization method for ancestral recombination graphs stored as tskit tree sequences
Interactive visualization method for ancestral recombination graphs stored as tskit tree sequences - GitHub - kitchensjn/tskit_arg_visualizer: Interactive visualization method for ancestral recomb...