Is there a rational way to predict stable engraftmet of a non-indigenous bacterial taxon in the context of an individual's gut microbiota?
In our latest preprint, @avcarrology tackles this fundamental question. @ISBNitinBaliga @thaasophobia @isbsci
https://www.biorxiv.org/content/10.1101/2023.04.28.538771v1
🧵
Prior work has looked at metagenomic features predictive of probiotic engraftment (
https://pubmed.ncbi.nlm.nih.gov/27693307/), ML approaches to predicting FMT donor strain engraftment (
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9499858/), or orthogonal niches that enable engraftment in the gut (
https://pubmed.ncbi.nlm.nih.gov/30110640/).

Stable Engraftment of Bifidobacterium longum AH1206 in the Human Gut Depends on Individualized Features of the Resident Microbiome - PubMed
Live bacteria (such as probiotics) have long been used to modulate gut microbiota and human physiology, but their colonization is mostly transient. Conceptual understanding of the ecological principles as they apply to exogenously introduced microbes in gut ecosystems is lacking. We find that, when …
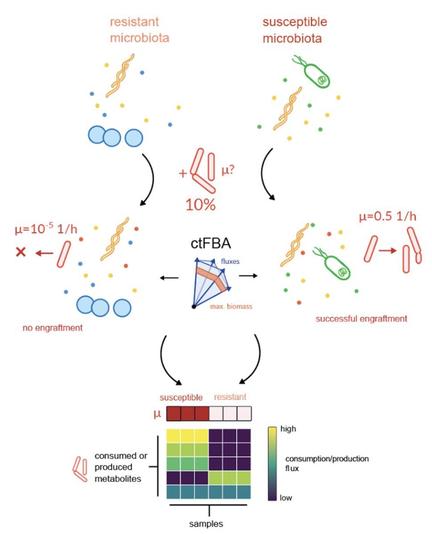
In this preprint, we use our MICOM platform to build microbial community-scale metabolic models (MCMMs) to predict personalized engraftment potential across >14,000 people.
We apply this approach to an important pathobiont known to colonize up to 30-40% of adults: C. difficile.
Basically, we 'inject' a non-indigenous taxon into a MCMM at a specified propagule pressure (e.g., 10% relative abundance), and then use its predicted growth rate as a proxy for engraftment/colonization probability within that individual (i.e., an engraftment risk index).
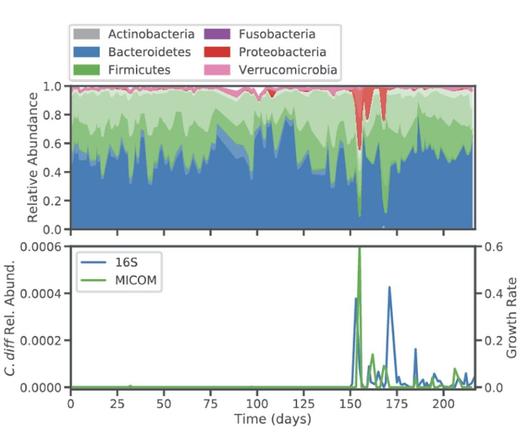
As an initial proof-of-principle for the accuracy of our MICOM/MCMM model, we leveraged time series data from a known asymptomatic C. difficile colonization event (following a bout of Salmonella food poisoning; https://www.nature.com/articles/s41564-020-0668-2).
The results were pretty good.

Diarrhoeal events can trigger long-term Clostridium difficile colonization with recurrent blooms - Nature Microbiology
Gastrointestinal disturbances, such as non-antibiotic-induced diarrhoea, promote susceptibility to colonization by Clostridium difficile, providing additional insights into antibiotic-independent triggers for this infection.
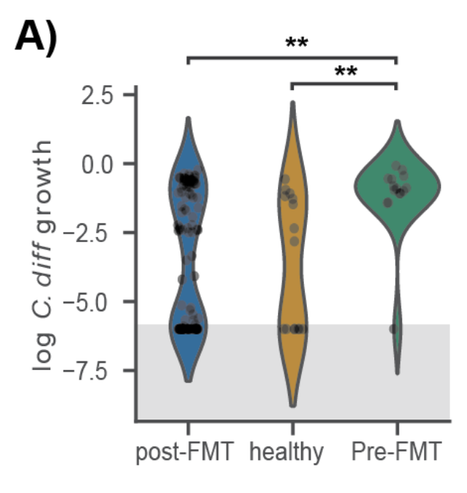
Next, we wanted to assess this engraftment risk index following an FMT treatment. We pulled down pre- and post-FMT sequencing data from
https://pubmed.ncbi.nlm.nih.gov/25825673/ and we saw that risk was high pre-FMT and lower post-FMT. However, many healthy individuals were susceptible to C. diff.

Dynamic changes in short- and long-term bacterial composition following fecal microbiota transplantation for recurrent Clostridium difficile infection - PubMed
Dynamic behavior is an intrinsic property of normal fecal microbiota and should be accounted for in comparing microbial communities among normal individuals and those with disease states. This also suggests that more frequent sample analyses are needed in order to properly assess success of FMT proc …
10^-6 was the lower limit of our solver accuracy, so any growth rate smaller than this number was effectively zero (i.e., see the grey shading in the prior plot).
Our metabolic model provides detailed mechanistic insights into what factors govern C. diff growth.
We looked at the top metabolites that C. diff consumed across models (i.e., its niche), & found that C. diff clustered into three growth niches: high-, moderate-, and no-growth.
The same exact genome-scale metabolic model of C. diff was used across these simulations, so these growth clusters show the plasticity of C. diff's metabolic niche across communities.
We see many metabolites known to promote C. diff growth from the literature, like succinate, ornithine, trehalose, and Stickland fermentation amino acids. We see other emergent metabolites important for growth as well, like fructose and cysteine/methionine.
MCMMs provide us with insight into potential competitors with C. diff, who import metabolites that it uses, like in the case of Phocaeicola and succinate (below). Even within the same taxon, these competitive interactions can vary a lot across individuals.
And MCMMs reveal taxa that cross-feed C. diff, like several of the butyrate-producing Clostridia, which appear to export succinate. Similarly, these cooperative interactions can vary a lot from person-to-person, even for the same bugs, highlighting the need for a systems approach
When we looked across 4 independent data sets, encompassing >14,000 individuals, we see that the three C. diff growth niches (import flux clusters) are highly consistent across populations.
If we embed these niches in the context of other commensal gut bacteria (grey clusters) in these models, we see that they are quite distinct from one another, relative to cross-taxon niche variation.
Previously, C. diff recurrence risk was observed to be higher in low-diversity microbiomes (
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5684761/). We saw this in our MCMMs too, & we found that engraftment risk also increased at the top-end of the diversity spectrum. Risk was lowest at intermediate diversity.

Identifying predictive features of Clostridium difficile infection recurrence before, during, and after primary antibiotic treatment
Colonization by the pathogen Clostridium difficile often occurs in the background of a disrupted microbial community. Identifying specific organisms conferring resistance to invasion by C. difficile is desirable because diagnostic and therapeutic strategies ...
Finally, as a proof-of-concept, we re-constructed a 6-member probiotic cocktail (i.e., a subset of VE303, with available metabolic models @VedantaBio;
https://jamanetwork.com/journals/jama/fullarticle/2803996), which was able to protect most MCMMs from C. diff invasion in silico.

Bacterial Consortium Therapy for Prevention of Recurrent C difficile Infection
This clinical trial compares the efficacy of 2 different doses of VE303, a defined bacterial consortium of 8 strains of commensal Clostridia, and placebo in preventing recurrence of Clostridioides difficile infection among adults at high risk.
However, there were clear responders and non-responders. The in silico efficacy of this probiotic cocktail was associated with the average growth rate of the probiotic taxa and/or with the average proximity of the probiotic taxa to the realized metabolic niche of C. diff.
And again, these models are very transparent, so we were able to look at how probiotics competed with C. diff for specific metabolites. We were also able to calculate the engraftment probability (growth rates) of the probiotoic strains themselves within each model.
We believe this work represents an advance in our ability to rationally predict personalized pathobiont colonization risk and assess efficacy of therapeutic probiotic interventions. Unlike more black-box machine learning approaches, MCMMs do not rely upon training data.
MCMMs provide transparent, mechanistically-grounded predictions that should be fairly robust across human populations. As our genome-scale metabolic model databases improve, these MCMMs will only grow more powerful.
This preprint is our first proof-of-concept for how MCMMs can be leveraged to predict engraftment potential. There is no reason why this same approach couldn't be applied to any other pathobiont or probiotic taxon in the human gut, or to more complex interventions, like FMTs.

Dynamic changes in short- and long-term bacterial composition following fecal microbiota transplantation for recurrent Clostridium difficile infection - PubMed
Dynamic behavior is an intrinsic property of normal fecal microbiota and should be accounted for in comparing microbial communities among normal individuals and those with disease states. This also suggests that more frequent sample analyses are needed in order to properly assess success of FMT proc …