Spatial-omics goes 3D🤗! Out at Cell 👉🏼 We developed DISCO-MS with @labs_mann, a spatial proteomics technology for specimens fully imaged in 3D. DISCO-MS is aided by robotics and enables the study of diseases at their early stages.
@cellpress https://www.cell.com/cell/fulltext/S0092-8674(22)01465-9🧵👇
Summary:
DISCO-MS is a spatial proteomics technology: it enables proteomics analysis of cleared tissues imaged in 3D. DISCO-MS is aided by AI and robotics and yields proteome similar to fresh or fixed samples.
Have questions? Add below👇 we will answer📖
Step 1:
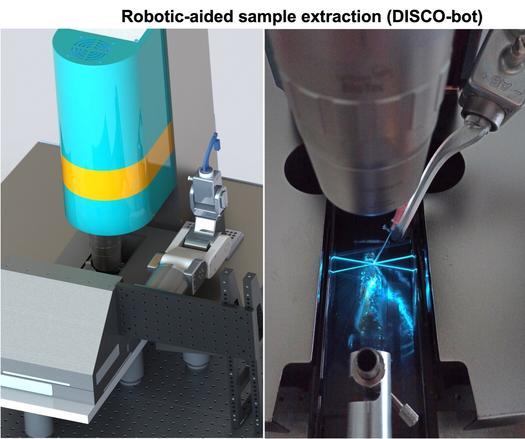
We start by rendering a mouse body or human organ optically transparent using robust clearing methods such as vDISCO or SHANEL. Then, after their cell-level light-sheet microscopy scan, we can visualize fluorescently-labeled structures in 3D as complete.
Step 2:
Next, we employ deep learning-based image analysis to identify regions associated with illnesses such as Alzheimer’s disease, brain injury, or coronary artery disease in an unbiased manner.
Step 3:
Small tissue regions are isolated with the help of a specialized robotic extraction system called DISCO-bot. Samples can currently be as small as 0.014 mm3. Smaller samples (down to 0.0005 mm3 or 60 cells) can be analyzed using laser capture microdissection.
Step 4:
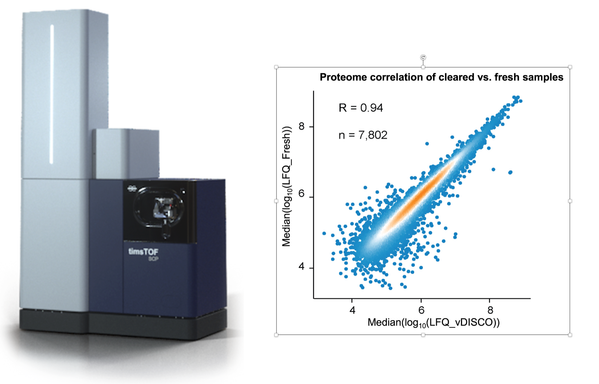
Next, the molecular composition of the tissue isolates is analyzed using high-sensitivity mass-spectrometry. The cellular proteome measured in cleared tissue is almost indistinguishable from the one obtained from fresh or fixed tissue controls.
Example application 1:
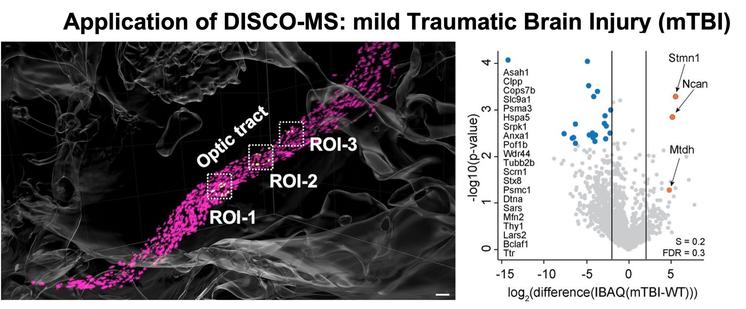
To illustrate the utility of DISCO-MS, we first characterized microglia activation after traumatic brain injury. We identified numerous proteins that were differentially regulated between equivalent regions along the optical tract in damaged and healthy brains.
Example application 2:
Our technology enables us to characterize subtle changes, which would be easily missed using standard methods. To this end, we analyzed the first A-beta plaques appearing in the 5xFAD Alzheimer’s disease mouse model from different brain regions.