https://doi.org/doi:10.1016/bs.apcsb.2025.10.031
https://pubmed.ncbi.nlm.nih.gov/41904007/
#Cytoskeleton #Cytoskeletal #CellDivision

Chromosomal instability (CIN), arising from errors in chromosome segregation during cell division, is a hallmark of cancer. Whilst CIN can result from several mitotic defects, the mechanisms that initiate CIN to drive tumourigenesis remain incompletely understood. Here, we show that loss of SMAD4 reprograms translation to induce CIN, resulting in tumour formation. Multi-omics analysis of tumourigenesis models driven by loss of SMAD4 complemented by functional studies demonstrate that loss of SMAD4 in pre-neoplastic cells leads to dysfunctional mitosis and an altered global translation landscape. We show that CDK11B is translationally downregulated in SMAD4 \ cells, and re-expression of the mitosis-specific isoform of this protein (CDK11B-p58) rescues the mitotic defects. Analysis of patient tumours reveals a strong correlation between markers of CIN and SMAD4 status, indicating the clinical relevance of this phenotype. Collectively, we reveal a previously unrecognised role for SMAD4 as a gatekeeper for CIN-mediated tumourigenesis via regulation of translation. ### Competing Interest Statement J.A. and M.W. are employees of Astex Pharmaceuticals. Astex Pharmaceuticals, UK National Health and Medical Research Council, #1182525

The biological significance of non-coding RNAs has been increasingly appreciated as their roles in various cellular processes are uncovered. However, single-cell transcriptomic profiling of human samples has focused primarily on protein-coding genes by targeting polyadenylated RNA transcripts, leaving the expression patterns of non-coding RNA underexplored. Here, we expand Tabula Sapiens to the non-coding transcriptome with single-cell and single-nucleus total RNA sequencing across 22 human organs and tissues. By simultaneously profiling both polyadenylated and non-polyadenylated transcripts, the resulting dataset enables joint analysis of the protein-coding and non-coding transcriptomes at single-cell and subcellular resolution. Using these data, we assessed the cell type specificity of non-coding genes and found that a greater proportion of non-coding genes are differentially expressed by single cell types compared to protein-coding genes. We then compared single-cell and single-nucleus data from the same samples to infer subcellular localization patterns, revealing cell type-dependent nuclear and cytoplasmic enrichment of specific non-coding RNAs. Next, we showed that tRNA repertoires are cell type-specific and that this specificity is not simply explained by differences in codon usage across cell types. Finally, we characterized dynamic expression patterns of non-coding RNAs across the cell cycle and senescence-associated cell states, identifying non-coding genes with putative roles in cell division and growth arrest. Our work establishes a resource for investigating the landscape of non-coding RNAs across a diverse set of human tissues and cell types. ### Competing Interest Statement The authors have declared no competing interest.

Multicellular organisms produced by sexual reproduction develop from single cells and the asymmetry of these cells can define the orientation of the earliest developmental axes. The haploid multicellular stage of the plant, Marchantia polymorpha, develops from a single cell - the spore - that divides asymmetrically, producing an apical germ cell that generates the plant body and a smaller basal cell that differentiates as an anchoring germ rhizoid cell. We show that the orientation of this asymmetric cell division is controlled by an external, environmental cue - blue light - that is perceived by the photoreceptor PHOTOTROPIN and signals in an NCH1-dependent manner. This defines core elements of the mechanism by which a directional environmental signal orients cell division, which in turn orients the first axis of symmetry. ### Competing Interest Statement Liam Dolan is a co-founder, shareholder, and board member of MoA Technology. Austrian Academy of Sciences, https://ror.org/03anc3s24, GMI Vienna European Research Council, DENOVO-P, 787613 European Molecular Biology Organization, https://ror.org/04wfr2810, Long-Term fellowship (2017)

Cell division in large embryos is coordinated by spatial waves of Cyclin B–Cdk1 activity that spread through the cytoplasm and affect cortical contractility. However, it is still unclear how cell size and localized activation near the nucleus shape these waves, and how the cytoplasmic signal is transmitted to the cortex. Here, we develop a reaction–diffusion model of Cyclin B–Cdk1 signaling in spherical cells with localized nuclear activation. We find that cytoplasmic waves have two distinct parts: an activation front that travels as a trigger wave, and a wave back that is controlled by inhibitory gradients in the cell cycle oscillator. Because these two parts are generated by different mechanisms, they can move at different speeds or even in opposite directions. This gives rise to different wave behaviors depending on nuclear size, nuclear position, and effective cell size. We then couple the Cdk1 signal to a cortical excitable network and show how cytoplasmic waveforms can regulate Rho–actin reactivation through inhibition of the RhoGEF Ect2. In this model, cortical patterns emerge mainly as downstream responses to cytoplasmic signaling, rather than as self-organized cortical waves. Overall, our results provide a mechanistic framework linking localized nuclear activation, cytoplasmic cell cycle waves, and cortical responses in large embryonic cells. ### Competing Interest Statement The authors have declared no competing interest.

To achieve a stereotypic lineage, each embryo of Caenorhabditis elegans follows an invariant cell differentiation process arising from a combination of cell polarisation, asymmetric or symmetric divisions, combined with intercellular signalling processes. This pattern of embryonic cell differentiation is driven by regulated segregation of molecules occurring at each cell division, including polarity proteins or cell fate determinants, transcription factors, p-granules and mRNAs. These distribution patterns are coupled with a robust spatio-temporal orchestration of cortical actin dynamics, which also plays a crucial role in these processes. However, compared to other molecular contents, how the actin per se is segregated from the first asymmetric division onward remains poorly understood. This study presents a thorough quantification of the intracellular distribution from the zygote to the 4-cell stage of key actors related to actin polymerisation: two nucleators (a formin and the Arp2/3 complex), a capping protein and E-cadherin. We additionally developed a novel method to assess actin polymerisation capacities from single blastomere extracts. We found that actin-related signatures arise at these early stages and that differential mechanisms of protein segregation and homeostasis occur, depending both on the cell pair and on the protein considered. Notably, if asymmetric divisions correlated with unequal partitioning of actin-related contents in a process linked with embryonic polarity, differences were revealed between AB daughter cells upon their separation. Taken together, these actin-related asymmetric distributions are adding a layer to the complexity of cell fate acquisition mechanisms in the early embryo. ### Competing Interest Statement The authors have declared no competing interest. Agence Nationale de la Recherche, ANR-19-CE13-0005-01
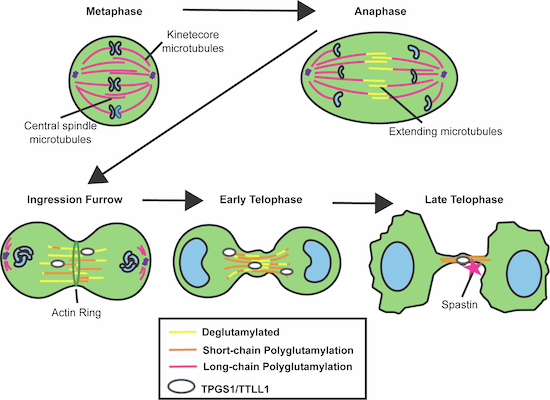
Microtubules perform a variety of cellular functions, including regulation of mitotic cell division, cilia formation, and neurite extension. Post-translational modifications controlled by the TTLL-family of enzymes confer a host of properties that affect microtubule dynamics and function. Specifically, polyglutamylation of tubulin C-terminal tails plays an important role in regulating microtubule dynamics and function within specific cellular contexts. In this study, we examined contributions from and potential regulators of polyglutamylation during mitosis, focusing on the microtubule remodeling that occurs in telophase once the mitotic spindle has completed chromosome separation. We demonstrate that the anaphase-to-telophase transition is accompanied by an increase in short-chain polyglutamylation of central spindle microtubules. We also show that TTLL1 and TPGS1, subunits of the tubulin polyglutamylation complex, are targeted to the intracellular bridge and midbody during cell progression through telophase. Finally, we demonstrate that loss of TPGS1 leads to defects in remodeling of the central spindle during telophase and impacts the cell’s ability to complete mitotic cell division.

Extrachromosomal DNA (ecDNA) is common in human cancers and is associated with poor clinical outcomes, yet how ecDNA-driven genetic heterogeneity is translated into functional heterogeneity remains unclear. Using single-cell multiomics sequencing and multiplexed IF-FISH, we show that asymmetric inheritance of ecDNA generates copy number heterogeneity that propagates to gene expression programs, including oncogenic signaling and cellular stress responses. Transgenerational live-cell lineage tracking directly shows that ecDNA heterogeneity arises within only a few cell divisions and modulates daughter cell division timing in a copy number-dependent manner, a property not observed for evenly inherited chromosomal amplicons. We identify an optimal middle ecDNA copy number range that maximizes proliferative fitness at baseline, while drug selection pressure induced by low-dose CHK1 inhibition selects for cells with a new optimal range at low ecDNA copy numbers. These low ecDNA copy number cells pre-exist in the population and can be generated de novo, driving copy number shifts promoting drug resistance. In vivo experiments further demonstrate that shifts toward ecDNA copy numbers that are optimal under the tumour microenvironment enhance tumourigenicity. Together, these findings establish ecDNA copy number plasticity as a central driver of tumour evolution. ### Competing Interest Statement P.S.M. is a co-founder of Boundless Bio and S1 Oncology. He has equity in both companies and consults, for which he is compensated. H.Y.C. is a co-founder of Accent Therapeutics, Boundless Bio, Cartography Biosciences, and Orbital Therapeutics and was an advisor to 10x Genomics, Arsenal Biosciences, Chroma Medicine, and Exai Bio until Dec 15, 2024. H.Y.C. is an employee and stockholder of Amgen as of Dec. 16, 2024.

In Gram-negative bacteria, co-translational insertion of membrane proteins into the plasma membrane may be coupled to ongoing transcription, a phenomenon known as transertion. Transertion results in a physical shift of the coding gene from the nucleoid towards the membrane and is one of the determinants of the shape of the nucleoid and placement of cell division in Gram-negative bacteria. In contrast, the existence of functional coupling of transcription and translation in Bacillus subtilis and potentially other Gram-positive bacteria has been questioned, suggesting that transertion may not happen. Here, by imaging vertically oriented B. subtilis cells, we show that the gene of a transmembrane protein changes its localization from inside the nucleoid to the plasma membrane upon induction of its transcription. Localization is restored to the nucleoid when induction of transcription has ceased. The shift of the gene towards the membrane is strictly dependent on transcription, its induction, translation and transmembrane nature of the coded protein. These results suggest that, at least, some principles of cellular regulation based on functional coupling of transcription and translation may be conserved between Gram-positive and Gram-negative bacteria. ### Competing Interest Statement The authors have declared no competing interest. Wellcome Trust, 217189/Z/19/Z BBSRC, BB/T017570/1