Materials: https://cosyne-tutorial-2025.github.io/
Materials: https://cosyne-tutorial-2025.github.io/
Which one are you at? #Cosyne2025
Last day of #Cosyne2025 before heading to the workshops! 🧠🗻🤓
Hope you enjoyed:
Neuroethology
From structure to function
Sensorimotor transformations
Circuit formation
Learning and plasticity
Systemic control of behavior
Dynamic cognitive computations
🧠🤓🧠🤓🧠
If you are at #COSYNE2025 today, I'm excited to suggest & self-promote work by amazing Ayesha Vermani. Poster session (Thu) 1-030:
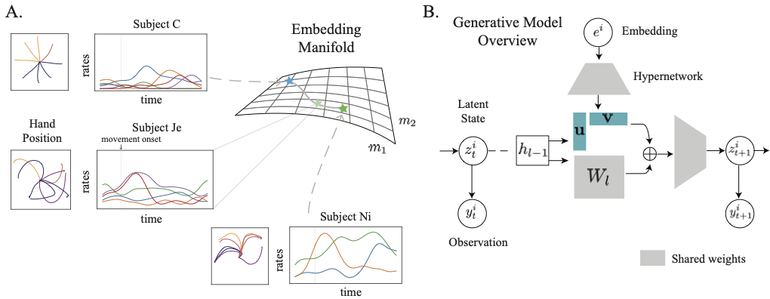
Meta-Dynamical State Space Models for Integrative Neural Data Analysis
Also ICLR'25 manuscript: https://openreview.net/forum?id=SRpq5OBpED
Check-out our poster at #cosyne2025 : "Robust Unsupervised Learning of Spike Patterns with Optimal Transport Theory" with Antoine Grimaldi, Matthieu Gilson, myself, @artipago.bsky.social Boris Sotomayor-Gomez, and Martin Vinck
https://laurentperrinet.github.io/publication/grimaldi-25-cosyne/
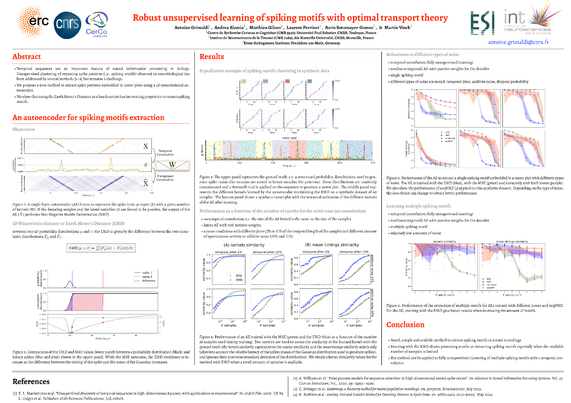
Robust Unsupervised Learning of Spike Patterns with Optimal Transport Theory | Next-generation neural computations
Temporal sequences are an important feature of neural information processing in biology. Neurons can fire a spike with millisecond precision, and, at the network level, repetitions of spatiotemporal spike patterns are observed in neurobiological data. However, methods for detecting precise temporal patterns in neural activity suffer from high computational complexity and poor robustness to noise, and quantitative detection of these repetitive patterns remains an open problem. Here, we propose a new method to extract spike patterns embedded in raster plots using a 1D convolutional autoencoder with the Earth Mover’s Distance (EMD) as a loss function. Importantly, the properties of the EMD make the method suitable for spike-based distributions, easy to compute, and robust to noise. Through gradient descent, the autoencoder is trained to minimize the EMD between the input and its reconstruction. We then expect the weight matrices to learn the repeating spike patterns present in the data. We validate our method on synthetically generated raster plots and compare its performance with an autoencoder trained using the Mean Squared Error (MSE) as a loss function. We show that the method using the EMD performs better at detecting the occurrence of the spike patterns, while the method using the MSE is better at capturing the underlying distributions used to generate the spikes. Finally, we propose to train the autoencoder iteratively by sequentially combining the EMD and the MSE losses. This sequential approach outperforms the widely used seqNMF method in terms of robustness to various types of noise, speed and stability. Overall, our method provides a novel approach to reliably extract repetitive temporal spike sequences, and can be readily generalized to other sequence detection applications.
👋 At #Cosyne2025?
Don't forget to check out work involving our researchers!
Use #cosyne2025 so we can 👀 what you're up to!!
If you are going to #cosyne2025 do check out Yijie Yin's poster 2-040 in Poster Session 2:
"Connectome Interpreter: a toolkit for efficient connectome exploration and hypothesis generation"
https://www.cosyne.org/poster-session-2
See also her software repository for the Connectome Interpreter:
https://github.com/YijieYin/connectome_interpreter
Going to #Cosyne2025? Check out our researchers’ posters and talks:
https://www.sainsburywellcome.org/web/content/swc-cosyne-2025
Wtih: Lars Rollik, Dammy Onih, Athena Akrami, Quentin Pajot-Moric, Margarida Perixia, Jasmine Reggiani, Laurence Freeman, Joanna Aloor, Yaara Lefler, @CampagnerDario, Joel Bauer, Svenja Nierwetberg, Tom George, Ali Haydaroglu, Jin Lee, Ann Duan, Neil Burgess, Lauren Bennett