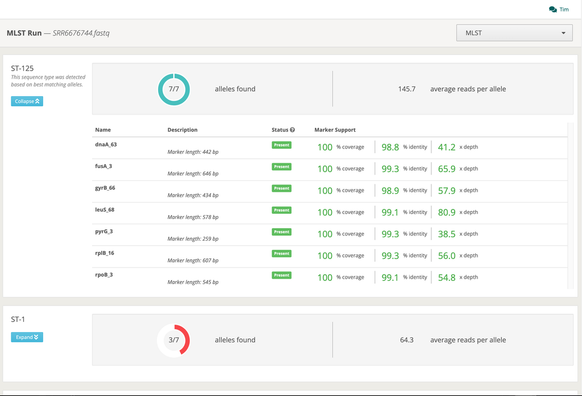
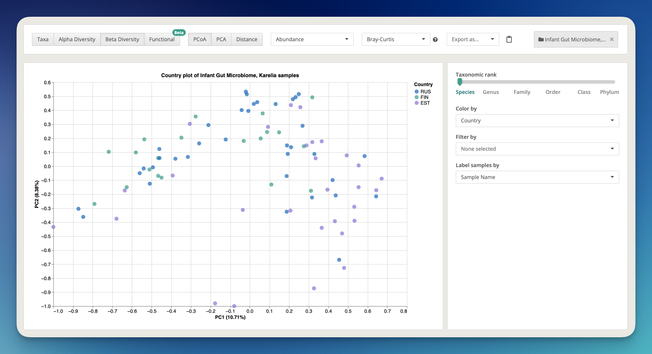
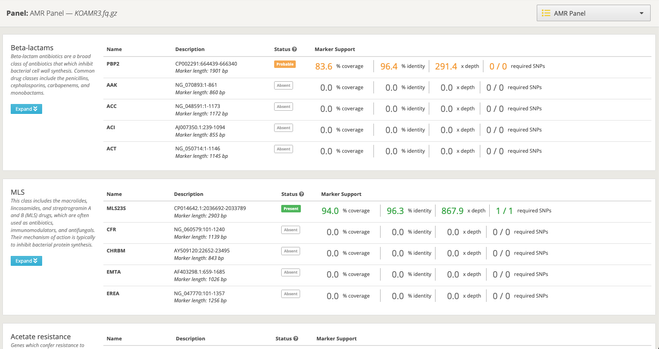
Major updates to MLST analysis at One Codex: we’ve expanded to 100+ species, added fungal support, and integrated long-read sequencing compatibility.
Read more about our revamped characterization workflow: https://www.onecodex.com/blog/2026/01/05/comprehensive-and-accurate-bacterial-isolate-characterization-using-multi-locus-sequence-typing/