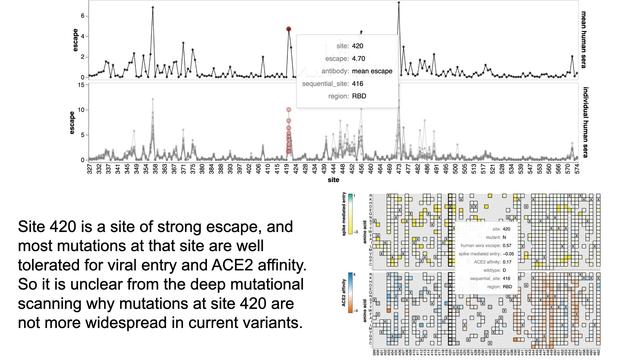
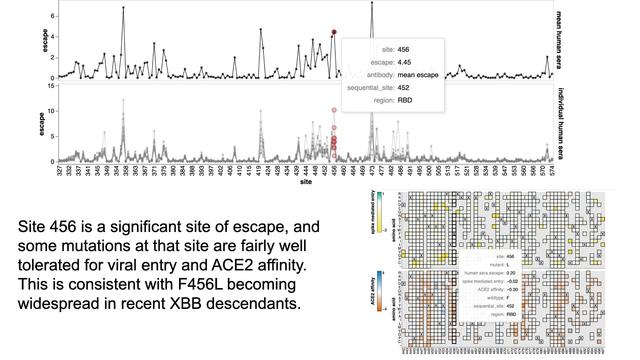
Here are data from new XBB.1.5 spike deep mutational scanning by Bernadeta Dadonaite in our group
Go to https://dms-vep.github.io/SARS-CoV-2_XBB.1.5_spike_DMS/htmls/summary_overlaid.html to see how mutations affect:
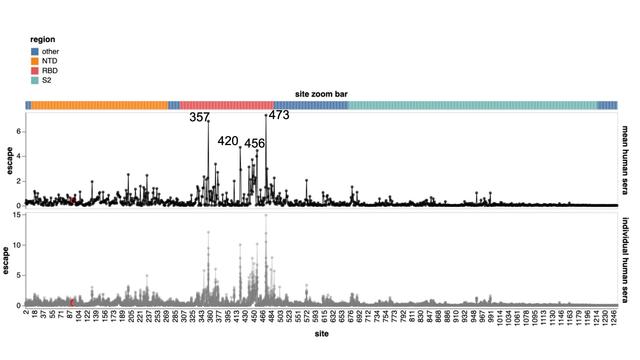
1⃣ Neutralization by human XBB breakthrough sera
2⃣ Spike-mediated pseudovirus entry in 293T-ACE2 cells
3⃣ Affinity for ACE2
It will be few weeks before we finish pre-print for study, but we wanted to share data now for those interested in SARS2 evolution
Measurements made using lentiviral deep mutational scanning: https://sciencedirect.com/science/article/pii/S0092867423001034