@me_the_fl00f Wouldn't it be RNA in this case?
@foxhkron Nope, DNA uses T (Thymin), RNA uses U (Uracil) then it would be AUG_AUUCAU...
The base idea was to hide a message in a CSS scanline based on a cyberpunk/biopunk/geopunk theme.
3 letters are one Codon represnts one amino acid. The hidden message is something like.
ATG → M (Methionin/Start)
ATT CAT GCT TGT AAA → I H A C K
ACT CAT GAG CGT GAG → T H E R E
TTT CAG CGT GAG → F Q R E
ATT GCT ATG → I A M
TAG → _ (Stop-Codon)
like M I HACK THEREFORE I AM based on Descartes Cogito, ergo sum. But hey, why not build encryption based on Gauß 
The base idea was to hide a message in a CSS scanline based on a cyberpunk/biopunk/geopunk theme.
3 letters are one Codon represnts one amino acid. The hidden message is something like.
ATG → M (Methionin/Start)
ATT CAT GCT TGT AAA → I H A C K
ACT CAT GAG CGT GAG → T H E R E
TTT CAG CGT GAG → F Q R E
ATT GCT ATG → I A M
TAG → _ (Stop-Codon)
like M I HACK THEREFORE I AM based on Descartes Cogito, ergo sum. But hey, why not build encryption based on Gauß

@me_the_fl00f Oh, I see. Only thought it was RNA because it’s only a single strand :D
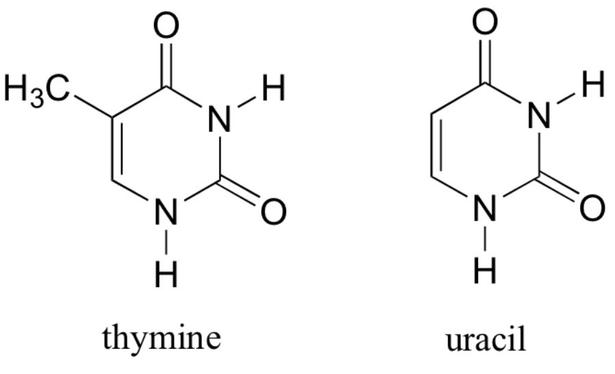
Uracil is structurally very close to thymine, but without the 5-methyl group.
Uracil is structurally very close to thymine, but without the 5-methyl group.
@foxhkron True, but you think in InfoSec structures, where a T is a T and a U is a U like a password will only match with the correct string or the correct hash of the string. In biologie first the T will become a U and then 1 triplet is the code for 1 amino acid, like as if you have 4 hashes matches for 1 string. In IT 1 wrong string decides about everything in biology it opens a whole rabbit hole of possibilities. Later or tomorrow I will publish a more detailed blog post. But in the meantime if you are bored https://thesai.org/Downloads/Volume12No10/Paper_21-A_Review_of_Modern_DNA_based_Steganography.pdf
@me_the_fl00f that’s interesting, thank you for the read :)
🐶 mood 💞 // [email protected]
Bound by paws, guided by spirit. Creating destiny through every high and low.
@foxhkron
The #Genetic Cipher: A #Steganographic Thought Experiment
Since you enjoyed the first DNA-splicing decode, let’s level up. Think of biological DNA not just as a blueprint, but as a multi-layered, encrypted filesystem.
1. The Biological Stack (Quick Primer)
To understand the „exploit“, you need to see how the „hardware“ reads the „code“:
- The Double Helix: DNA consists of two strands. They are antiparallel (running in opposite directions, 5′ → 3′ vs. 3′ → 5′ ) and complementary (A pairs with T, C pairs with G).
- Transcription: The cell doesn't read the whole „disk“. It transcribes DNA into mRNA.
- Splicing: Here’s the InfoSec twist: A gene contains Exons (coding data) and Introns (non-coding „junk“ data). During splicing, the Introns are removed, and only Exons are joined to form the final „executable“ protein.
- The Codon Table: 3 bases = 1 amino acid. But it’s a degenerate code (multiple „opcodes“ can result in the same output), which is perfect for hiding noise.
2. The „Gauss-Intron“ Proposal
If we wanted to hide a message that even a sophisticated LLM or pattern-scanner wouldn't flag, we’d use the Introns as a steganographic layer:
- Antiparallel Mirroring: We hide the message on the Template Strand (the „backside“ of the DNA). To find it, you’d have to flip the string and calculate the complement -- basically a biological „reverse-text“ cipher.
- The Gauss Mask: Instead of hiding text in the Introns (which would look like suspicious „high-entropy“ data), we hide the metadata (start/stop triggers) in their statistical distribution.
- The Logic: If the lengths of the Introns follow a perfect Gaussian Distribution (Bell Curve), any automated bio-scanner would mark it as „natural variation“ or „evolutionary noise“.
- The Language Key: To avoid translation errors across different „compilers“, we use Latin (a dead, rigid language) for the Exons. It acts as a „Proof of Concept“ that the decoded text isn't a fluke.
The Challenge:
Imagine a sequence where the Exons tell a boring story in Latin, but the Introns -- invisible to the final protein -- dictate the decryption key for the reverse-complement strand based on their relative positions on a Gaussian curve.
„Nature doesn't just hide its secrets in the code, but in the way the code is edited before execution“.
Example:
5'--ATG_gtacagttgc_GGT_agctagctagctagct_GTG_gtacagttgc_GTG_tag--3'
Introns lower case, Exons upper case, slipcing result ATG GGT GTG GTG TAG
ATG → M (Methionin/Start)
GGT → G
GTG → V
GTG → V
TAG → _ (Stop)
Result: "MGVV" Magna Vis.
Count the introns, maybe you get 10 - 16 - 10 (you remember Bell Curve). Or is it a trigger for the message at the 3' → 5' Strang, who knows.
Keep in mind, first comes DNA (T) then RNA (U) with which you can use the Code-Sonne on Wikipedia.
The #Genetic Cipher: A #Steganographic Thought Experiment
Since you enjoyed the first DNA-splicing decode, let’s level up. Think of biological DNA not just as a blueprint, but as a multi-layered, encrypted filesystem.
1. The Biological Stack (Quick Primer)
To understand the „exploit“, you need to see how the „hardware“ reads the „code“:
- The Double Helix: DNA consists of two strands. They are antiparallel (running in opposite directions, 5′ → 3′ vs. 3′ → 5′ ) and complementary (A pairs with T, C pairs with G).
- Transcription: The cell doesn't read the whole „disk“. It transcribes DNA into mRNA.
- Splicing: Here’s the InfoSec twist: A gene contains Exons (coding data) and Introns (non-coding „junk“ data). During splicing, the Introns are removed, and only Exons are joined to form the final „executable“ protein.
- The Codon Table: 3 bases = 1 amino acid. But it’s a degenerate code (multiple „opcodes“ can result in the same output), which is perfect for hiding noise.
2. The „Gauss-Intron“ Proposal
If we wanted to hide a message that even a sophisticated LLM or pattern-scanner wouldn't flag, we’d use the Introns as a steganographic layer:
- Antiparallel Mirroring: We hide the message on the Template Strand (the „backside“ of the DNA). To find it, you’d have to flip the string and calculate the complement -- basically a biological „reverse-text“ cipher.
- The Gauss Mask: Instead of hiding text in the Introns (which would look like suspicious „high-entropy“ data), we hide the metadata (start/stop triggers) in their statistical distribution.
- The Logic: If the lengths of the Introns follow a perfect Gaussian Distribution (Bell Curve), any automated bio-scanner would mark it as „natural variation“ or „evolutionary noise“.
- The Language Key: To avoid translation errors across different „compilers“, we use Latin (a dead, rigid language) for the Exons. It acts as a „Proof of Concept“ that the decoded text isn't a fluke.
The Challenge:
Imagine a sequence where the Exons tell a boring story in Latin, but the Introns -- invisible to the final protein -- dictate the decryption key for the reverse-complement strand based on their relative positions on a Gaussian curve.
„Nature doesn't just hide its secrets in the code, but in the way the code is edited before execution“.
Example:
5'--ATG_gtacagttgc_GGT_agctagctagctagct_GTG_gtacagttgc_GTG_tag--3'
Introns lower case, Exons upper case, slipcing result ATG GGT GTG GTG TAG
ATG → M (Methionin/Start)
GGT → G
GTG → V
GTG → V
TAG → _ (Stop)
Result: "MGVV" Magna Vis.
Count the introns, maybe you get 10 - 16 - 10 (you remember Bell Curve). Or is it a trigger for the message at the 3' → 5' Strang, who knows.
Keep in mind, first comes DNA (T) then RNA (U) with which you can use the Code-Sonne on Wikipedia.