1/5
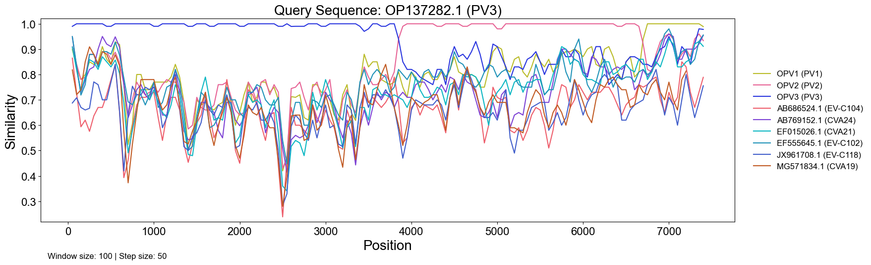
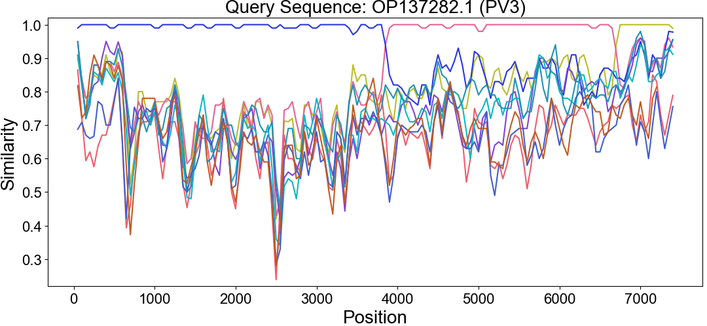
Introducing SimPlot-CL! 🧬
Recombination plays an important role in viral evolution. Similarity plots are a great way to visualize recombination patterns, but generating them across many genomes can be cumbersome
(Work with my student, Keno Strotjohann!)
📂 github.com/hodcroftlab/simplot-cl