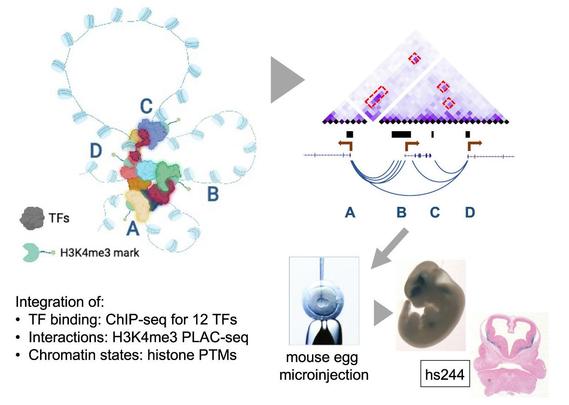
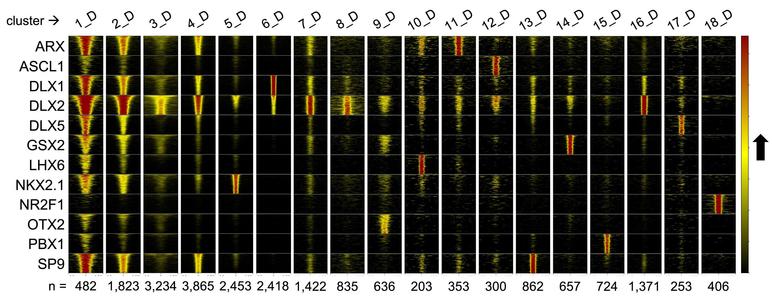
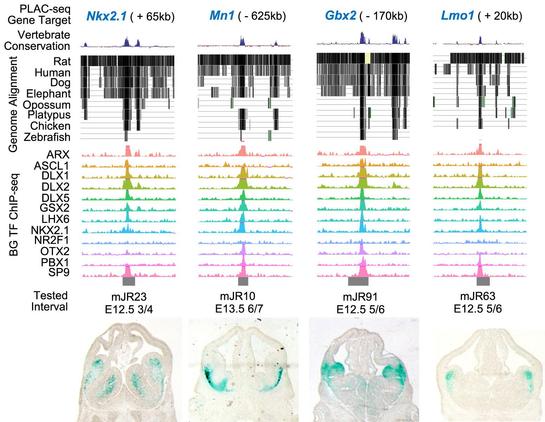
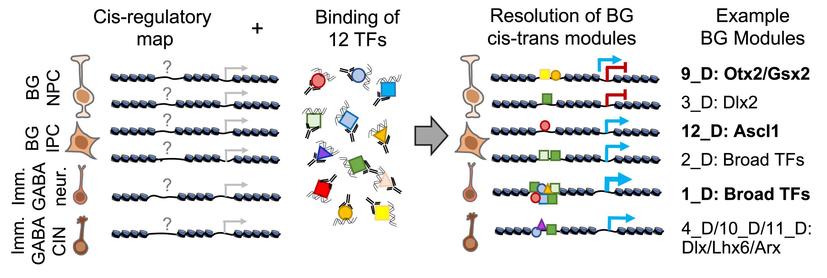
🧵 Paper alert: How do different transcription factors work together to help form GABAergic neurons in the brain?
The work was led by Rinaldo Catta-Preta, Susan Lindtner, John Rubenstein, and Alex Nord (we were a minor part of it) - time for a thread nonetheless! 1/14
https://authors.elsevier.com/sd/article/S1534-5807(24)00603-8