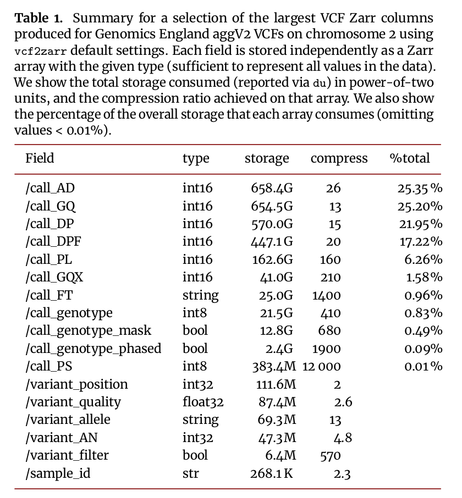
Excited to share my latest preprint (with a stellar band of collaborators), where we map the VCF data model into an efficient, cloud-native storage format. Thread follows:
https://www.biorxiv.org/content/10.1101/2024.06.11.598241v1
https://www.biorxiv.org/content/10.1101/2024.06.11.598241v1