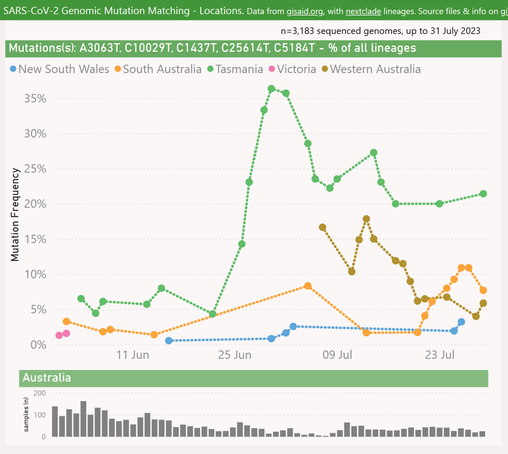
Here's the latest variant picture for Tasmania, Australia.
The "Deltacron" XAY.* variant (33%) has put on a surprising sudden surge from late July onwards. This might explain the rise in cases and hospitalisations during that period.
XAY is a recombinant of Omicron BA.2 and Delta AY.45. XAY.* has been slowly evolving: XAY.1 added the Spike D253G mutation, then XAY.1.1 added Spike R346T and ORF1a K2108N. XAY.1.1.1 added Spike D1153Y and finally child lineage GL.1 added Spike D420N. GL.1 is the only sublineage present in the recent data from Tasmania.
🧵