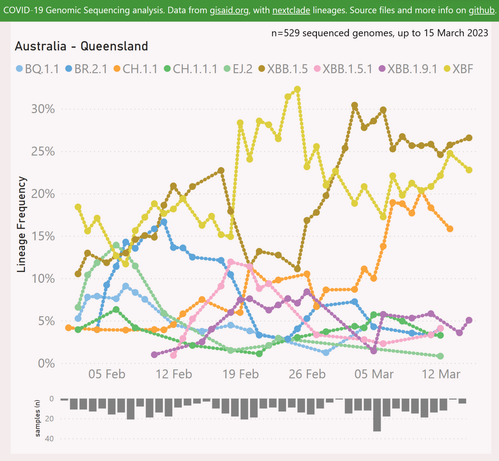
Here's the latest variant picture for Queensland.
XBB.1.5 "Kraken" (27%) seems dominant, although XBF "Bythos" (23%) is putting up stubborn resistance.
There's an unexpected surge from CH.1.1 "Orthrus" (16%). While this seems a robust variant (causing significant waves earlier in the UK, Hong Kong and NZ), elsehwere it has been suppressed by XBB.1.5 and XBF. Perhaps a new sub-variant has arisen? This would be nsurprising, given the lack of masking in hospitals in QLD. That creates an environment where immunocompromised folks are forced into multiple infections - the perfect breeding ground for new variants.
XBB.1.9.1 "Hyperion" (5%) is relatively low, compared to other states.
The sample volumes seem representative up to 13 March.
Interactive genomic sequencing dataviz, code, acknowledgements and more info here:
https://github.com/Mike-Honey/covid-19-genomes#readme
#COVID19 #Australia #QLD #Brisbane #XBB_1_5 #Kraken @auscovid19