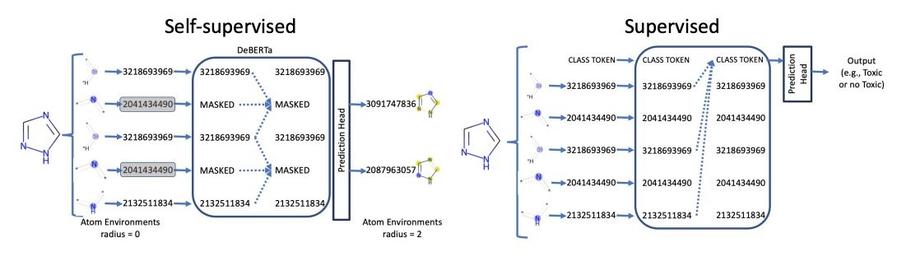
Two-stage pretraining for chemicals:
1. Masked language model
2. Predict chemical properties
@omendezlucio Nicolaou, @bertonearnshaw
Interpolated polynomial multiple zeta values of fixed weight, depth, and height
We define the interpolated polynomial multiple zeta values as a generalization of all of multiple zeta values, multiple zeta-star values, interpolated multiple zeta values, symmetric multiple zeta values, and polynomial multiple zeta values. We then compute the generating function of the sum of interpolated polynomial multiple zeta values of fixed weight, depth, and height.