This landmark #ReviewedPreprint represents “the most significant breakthrough in membrane and secretory biogenesis in recent years.”
‘Structural analysis of the dynamic ribosome-translocon complex’
https://elifesciences.org/reviewed-preprints/95814v2?utm_source=mastodon&utm_medium=social&utm_campaign=organic
Structural analysis of the dynamic ribosome-translocon complex
This landmark #ReviewedPreprint represents “the most significant breakthrough in membrane and secretory biogenesis in recent years.”
‘Structural analysis of the dynamic ribosome-translocon complex’
https://elifesciences.org/reviewed-preprints/95814v2?utm_source=mastodon&utm_medium=social&utm_campaign=organic
Structural analysis of the dynamic ribosome-translocon complex
What’s it actually like to publish a #ReviewedPreprint with eLife?
We asked author Chunxiao Li, a palaeontologist at the University of Chinese Academy of Sciences. Read about her experience.
https://elifesciences.org/inside-elife/f87ed206/author-experience-innovative-and-intriguing?utm_source=mastodon&utm_medium=social&utm_campaign=organic

Author Experience: “innovative and intriguing”
Chunxiao Li, postdoctoral researcher in vertebrate paleontology, University of Chinese Academy of Sciences, Beijing, China, shares her experience of publishing a Reviewed Preprint with eLife.
Most species have two sexes. The protist Tetrahymena thermophila has seven.
This fundamental #ReviewedPreprint reveals how it recognises self and non-self mating-types with a novel protein complex. https://elifesciences.org/reviewed-preprints/93770?utm_source=mastodon&utm_medium=social&utm_campaign=organic
A seven-sex species recognizes self and non-self mating-type via a novel protein complex
Biomineralisation appears to have evolved multiple times using different materials. This
#ReviewedPreprint reveals how ROCK, an actomyosin remodelling protein, controls the process in sea urchin larvae.
https://elifesciences.org/reviewed-preprints/89080?utm_source=mastodon&utm_medium=social&utm_campaign=organicROCK and the actomyosin network control biomineral growth and morphology during sea urchin skeletogenesis
New #ReviewedPreprint presents a method to fluorescently label peptide MHC complexes on live dendritic cells, allowing for a better understanding of antigen-specific T cell interactions.
Public reviews by @eLife.
https://sciety.org/articles/activity/10.21203/rs.3.rs-3193191/v2?utm_source=mastodon&utm_medium=social&utm_campaign=sci_mastodon_2024_15
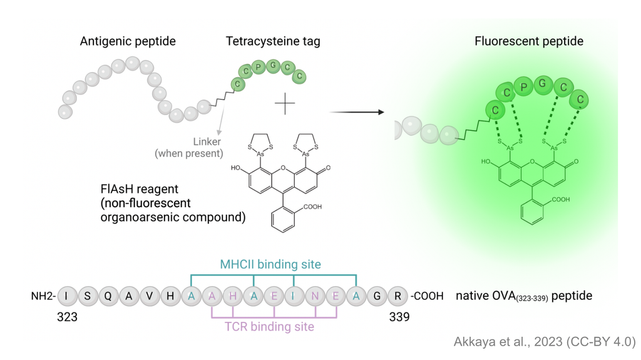

Illuminating T cell-dendritic cell interactions in vivo by FlAsHing antigens
Delineating the complex network of interactions between antigen-specific T cells and antigen presenting cells (APCs) is crucial for effective precision therapies against cancer, chronic infections, and autoimmunity. However, the existing arsenal for examining antigen-specific T cell interactions is restricted to a select few antigen-T cell receptor pairs, with limited in situ utility. This lack of versatility is largely due to the disruptive effects of reagents on the immune synapse, which hinder real-time monitoring of antigen-specific interactions. To address this limitation, we have developed a novel and versatile immune monitoring strategy by adding a short cysteine-rich tag to antigenic peptides that emits fluorescence upon binding to thiol-reactive biarsenical hairpin compounds. Our findings demonstrate the specificity and durability of the novel antigen-targeting probes during dynamic immune monitoring in vitro and in vivo. This strategy opens new avenues for biological validation of T-cell receptors with newly identified epitopes by revealing the behavior of previously unrecognized antigen-receptor pairs, expanding our understanding of T cell responses.
Analysing T cell motion in different tissue environments: many parameters of T cell motion are similar across tissues, with the exception of parameters related to confinement.
#ReviewedPreprint with public reviews by @eLife.
https://sciety.org/articles/activity/10.1101/2022.11.17.516891?utm_source=mastodon&utm_medium=social&utm_campaign=sci_mastodon_2024_13

Quantitative analyses of T cell motion in tissue reveals factors driving T cell search in tissues
T cells are required to clear infection, and T cell motion plays a role in how quickly a T cell finds its target, from initial naive T cell activation by a dendritic cell to interaction with target cells in infected tissue. To better understand how different tissue environments affect T cell motility, we compared multiple features of T cell motion including speed, persistence, turning angle, directionality, and confinement of T cells moving in multiple murine tissues using microscopy. We quantitatively analyzed naive T cell motility within the lymph node and compared motility parameters with activated CD8 T cells moving within the villi of small intestine and lung under different activation conditions. Our motility analysis found that while the speeds and the overall displacement of T cells vary within all tissues analyzed, T cells in all tissues tended to persist at the same speed. Interestingly, we found that T cells in the lung show a marked population of T cells turning at close to 180
o
, while T cells in lymph nodes and villi do not exhibit this “reversing” movement. T cells in the lung also showed significantly decreased meandering ratios and increased confinement compared to T cells in lymph nodes and villi. These differences in motility patterns led to a decrease in the total volume scanned by T cells in lung compared to T cells in lymph node and villi. These results suggest that the tissue environment in which T cells move can impact the type of motility and ultimately, the efficiency of T cell search for target cells within specialized tissues such as the lung.
The origins of RNA viruses can be hard to trace. This
#ReviewedPreprint considers a more recent origin for coronaviruses and suggests their diversification occurred mainly through host switching rather than diversification with hosts.
https://elifesciences.org/reviewed-preprints/91745?utm_source=mastodon&utm_medium=social&utm_campaign=organicRecent evolutionary origin and localized diversity hotspots of mammalian coronaviruses
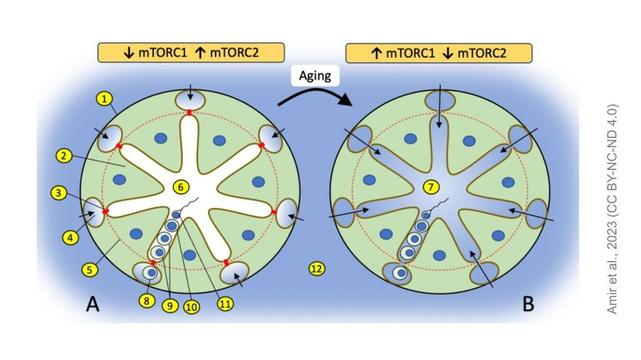

Mechanistic target of rapamycin (mTOR) pathway in Sertoli cells regulates age-dependent changes in sperm DNA methylation
Over the past several decades, a trend toward delayed childbirth has led to increases in parental age at the time of conception. Sperm epigenome undergoes age-dependent changes increasing risks of adverse conditions in offspring conceived by fathers of advanced age. Animal experiments also demonstrate that epigenetic transfer of maladaptive information of advanced age fathers is associated with reduced offspring health including reduced life span. The mechanism(s) linking paternal age with epigenetic changes in sperm remain unknown. The sperm epigenome is shaped in a compartment protected by the blood-testes barrier (BTB) known to deteriorate with age. Permeability of the BTB is regulated by the balance of two mTOR complexes in Sertoli cells. We hypothesized that this balance is also responsible for age-dependent changes in the sperm epigenome. Experiments with transgenic mice demonstrate that the shift of the balance in favor of mTOR complex two rejuvenates sperm DNA-methylome while the shift in favor of mTOR complex one accelerates aging of the sperm DNA-methylome and results in a reproductive phenotype concordant with older age. These results demonstrate for the first time that the balance of mTOR complexes in Sertoli cells regulates the rate of sperm epigenetic aging. Thus, mTOR pathway in Sertoli cells may be used as novel target of therapeutic interventions to rejuvenate the sperm epigenome in advanced-age fathers.
What controls the repair of photosystem II, a key process in maintaining and optimising photosynthesis?
Tryptophan oxidation plays a critical role, new #ReviewedPreprint with public reviews by @eLife suggests.
https://sciety.org/articles/activity/10.1101/2023.04.20.537607?utm_source=mastodon&utm_medium=social&utm_campaign=sci_mastodon_2023_209

Characterization of tryptophan oxidation affecting D1 degradation by FtsH in the photosystem II quality control of chloroplasts
Photosynthesis is one of the most important reactions for sustaining our environment. Photosystem II (PSII) is the initial site of photosynthetic electron transfer by water oxidation. Light in excess, however, causes the simultaneous production of reactive oxygen species (ROS), leading to photo-oxidative damage in PSII. To maintain photosynthetic activity, the PSII reaction center protein D1, which is the primary target of unavoidable photo-oxidative damage, is efficiently degraded by FtsH protease. In PSII subunits, photo-oxidative modifications of several amino acids such as Trp have been indeed documented, whereas the linkage between such modifications and D1 degradation remains elusive. Here, we show that an oxidative post-translational modification of Trp residue at the N-terminal tail of D1 is correlated with D1 degradation by FtsH during high-light stress. We revealed that
Arabidopsis
mutant lacking FtsH2 had increased levels of oxidative Trp residues in D1, among which an N-terminal Trp-14 was distinctively localized in the stromal side. Further characterization of Trp-14 using chloroplast transformation in
Chlamydomonas
indicated that substitution of D1 Trp-14 to Phe, mimicking Trp oxidation enhanced FtsH-mediated D1 degradation under high light, although the substitution did not affect protein stability and PSII activity. Molecular dynamics simulation of PSII implies that both Trp-14 oxidation and Phe substitution cause fluctuation of D1 N-terminal tail. Furthermore, Trp-14 to Phe modification appeared to have an additive effect in the interaction between FtsH and PSII core in vivo. Together, our results suggest that the Trp oxidation at its N-terminus of D1 may be one of the key oxidations in the PSII repair, leading to processive degradation by FtsH.






