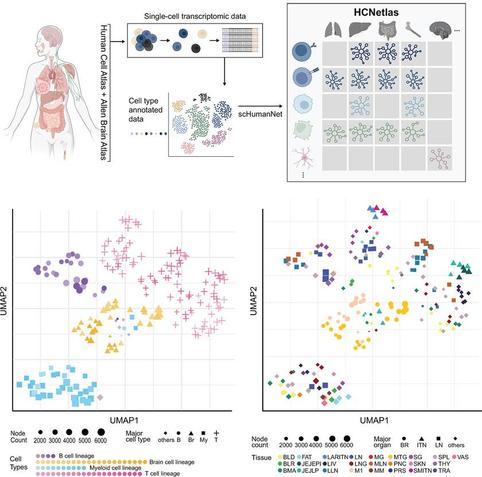
HCNetlas: A reference database of human cell type-specific gene networks to aid disease genetic analyses
Alterations of gene regulatory networks in specific cell types can cause disease. These authors create HCNetlas (Human Cell Network Atlas), a compilation of cell-type-specific gene networks across a range of healthy tissues that can be used to uncover associations between disease genes and specific cell types.